| Attribute | Type | Description |
|---|---|---|
| stime | Numeric | survival or follow-up time, in days |
| status | Numeric | dead or alive/censored; 0 (alive), 1 (dead) |
| treat | Nominal | treatment, either standard or test (chemotherapy); 1 (standard), 2 (test) |
| age | Numeric | patient age in years |
| Karn | Numeric | Karnofsky score of patient performance, on scale of 0 to 100 |
| diag | Numeric | patient time since diagnosis, measured at time of entry to trial, in months |
| cell | Nominal | one of four cell types; 1 (squamous), 2 (small cell), 3 (adeno), 4 (large) |
| prior | Nominal | denotes prior therapy; 10 (yes), 0 (no) |
Cox Proportional Hazards Model
Edwin Alvarado, Darnell Thomas, and Michael Ho
Overview
What is the Cox Proportional Hazards Model?
Used to evaluate the association between survival time of a patient and one or more risk factors
Known for its ability to handle censored data
Can account for “realistic” outcomes of observations (i.e., the patient surviving)
Defining the Statistical Model
The composition of the model is as follows:
\[ h_i(t) = h_0(t) \times exp(\beta_1X_{1i} + \beta_2X_{2i} + ... + \beta_kX_{ki}) \]
Where \(H_0 (t)\) is considered to be the baseline hazard function of the model. \(\beta_k\) are model parameters.
The Hazard Ratio
Consider the following example:
\[ HR = {h_{pre}(t) \over h_{post}(t)} = {h_0(t)\times exp(0.0177 \times X_{pre}) \over h_0(t) \times exp(0.0177 \times X_{post})} \] \[ = {exp(0.0177 \times 1) \over exp(0.0177 \times 0)} = 1.018\] Similar to Odds Ratio in that:
- HR > 1 = Positive Effect
- HR < 1 = Negative Effect
- HR = 1 = No Effect
Applications and Limitations
Often used in medical field, e.g. clinical trials
While commonly used in medical research, can be applied to other fields, such as finance or social sciences
Originally form of the model, could not account for time-dependent variable effects
- Proportional Hazard Assumption
Case Study: VA Lung Cancer Trial
Goals
- Determine if treatment used in trial has positive effect on survival time
- Determine if any other variables have significant effect on survival time
- Properly test assumptions and fit the Cox Proportional Hazards Model
Case Study: VA Lung Cancer Trial
| stime | status | treat | age | Karn | diag.time | cell | prior |
|---|---|---|---|---|---|---|---|
| 72 | 1 | 1 | 69 | 60 | 7 | 1 | 0 |
| 411 | 1 | 1 | 64 | 70 | 5 | 1 | 10 |
| 228 | 1 | 1 | 38 | 60 | 3 | 1 | 0 |
| 126 | 1 | 1 | 63 | 60 | 9 | 1 | 10 |
| 118 | 1 | 1 | 65 | 70 | 11 | 1 | 10 |
| 10 | 1 | 1 | 49 | 20 | 5 | 1 | 0 |
- 137 Observations of patients with inoperable lung cancer
- Includes Censored Data
- Per the National Cancer Institute, the Karnofsky score is standard way to measure how well a patient can perform ordinary tasks. A higher score indicates a higher ability to do these tasks
Summary Statistics
| stime | age | Karn | diag.time | |
|---|---|---|---|---|
| Min. : 1.0 | Min. :34.00 | Min. :10.00 | Min. : 1.000 | |
| 1st Qu.: 25.0 | 1st Qu.:51.00 | 1st Qu.:40.00 | 1st Qu.: 3.000 | |
| Median : 80.0 | Median :62.00 | Median :60.00 | Median : 5.000 | |
| Mean :121.6 | Mean :58.31 | Mean :58.57 | Mean : 8.774 | |
| 3rd Qu.:144.0 | 3rd Qu.:66.00 | 3rd Qu.:75.00 | 3rd Qu.:11.000 | |
| Max. :999.0 | Max. :81.00 | Max. :99.00 | Max. :87.000 |
| status | treat | cell | prior | |
|---|---|---|---|---|
| 0: 9 | 1:69 | 1:35 | 0 :97 | |
| 1:128 | 2:68 | 2:48 | 10:40 | |
| NA | NA | 3:27 | NA | |
| NA | NA | 4:27 | NA |
Data Visualization
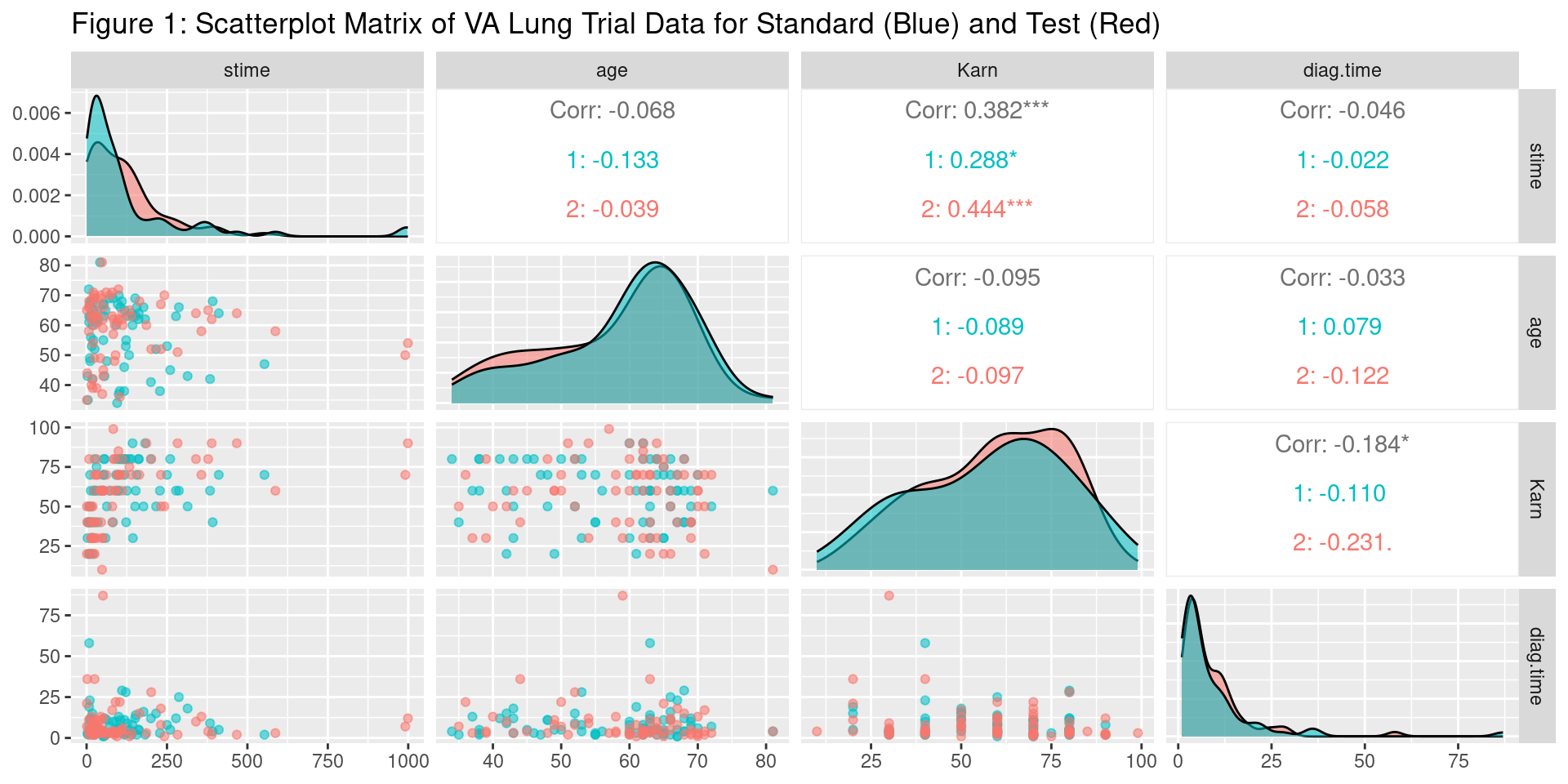
Distributions are roughly similar between standard and test treatment groups
Karnofsky and survival time seem positively correlated
#Create a scatter plot matrix using the GGally package
library(GGally)
library(tidyverse)
va_subset <- VA[, c("stime", "age", "Karn", "diag.time", "treat")]
ggpairs(va_subset, title = "Figure 1: Scatterplot Matrix of VA Lung Trial Data for Standard (Blue) and Test (Red)",
columns = c("stime", "age", "Karn", "diag.time"), ggplot2::aes(color = treat),
lower = list(continuous = "points", combo = "dot_no_facet", mapping = ggplot2::aes(color=treat, alpha = 0.8)),
diag = list(continuous = wrap("densityDiag"), mapping = ggplot2::aes(color = treat, alpha = 0.1))) +
scale_color_manual(values = c("#00BFC4", "#F8766D"))Statistical Model
Recall the composition:
\[ h_i(t) = h_0(t) \times exp(\beta_1X_{1i} + \beta_2X_{2i} + ... + \beta_kX_{ki}) \]
Model in R
Cox PH model for a single variable (treat)
# Import packages
library(survival)
# Fit Cox PH Model for treatment only
Y = Surv(VA$stime, VA$status)
coxph(Y ~ treat, data=VA)Call:
coxph(formula = Y ~ treat, data = VA)
coef exp(coef) se(coef) z p
treat2 0.01774 1.01790 0.18066 0.098 0.922
Likelihood ratio test=0.01 on 1 df, p=0.9218
n= 137, number of events= 128 Call:
coxph(formula = Y ~ treat, data = VA)
n= 137, number of events= 128
coef exp(coef) se(coef) z Pr(>|z|)
treat2 0.01774 1.01790 0.18066 0.098 0.922
exp(coef) exp(-coef) lower .95 upper .95
treat2 1.018 0.9824 0.7144 1.45
Concordance= 0.525 (se = 0.026 )
Likelihood ratio test= 0.01 on 1 df, p=0.9
Wald test = 0.01 on 1 df, p=0.9
Score (logrank) test = 0.01 on 1 df, p=0.9Proportional Hazard Assumption
The proportional hazard assumption is means that hazard ratio (HR) is constant over time.
Variables and regression coefficients are time-independent (no change over course of study).
Methods of checking PH assumption
Schoenfeld Residuals vs Time Plot
Statistical test of Schoenfeld Residuals
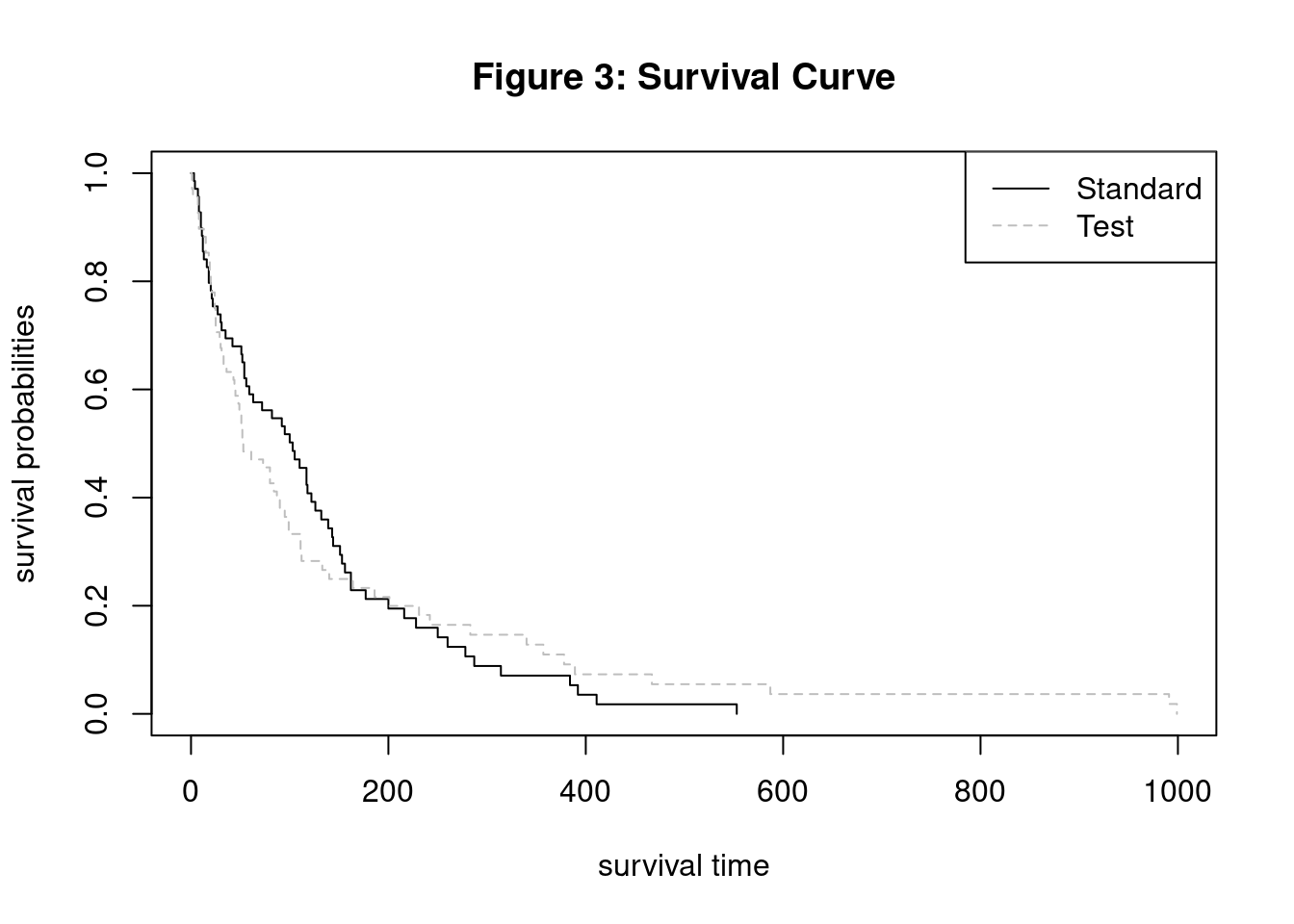
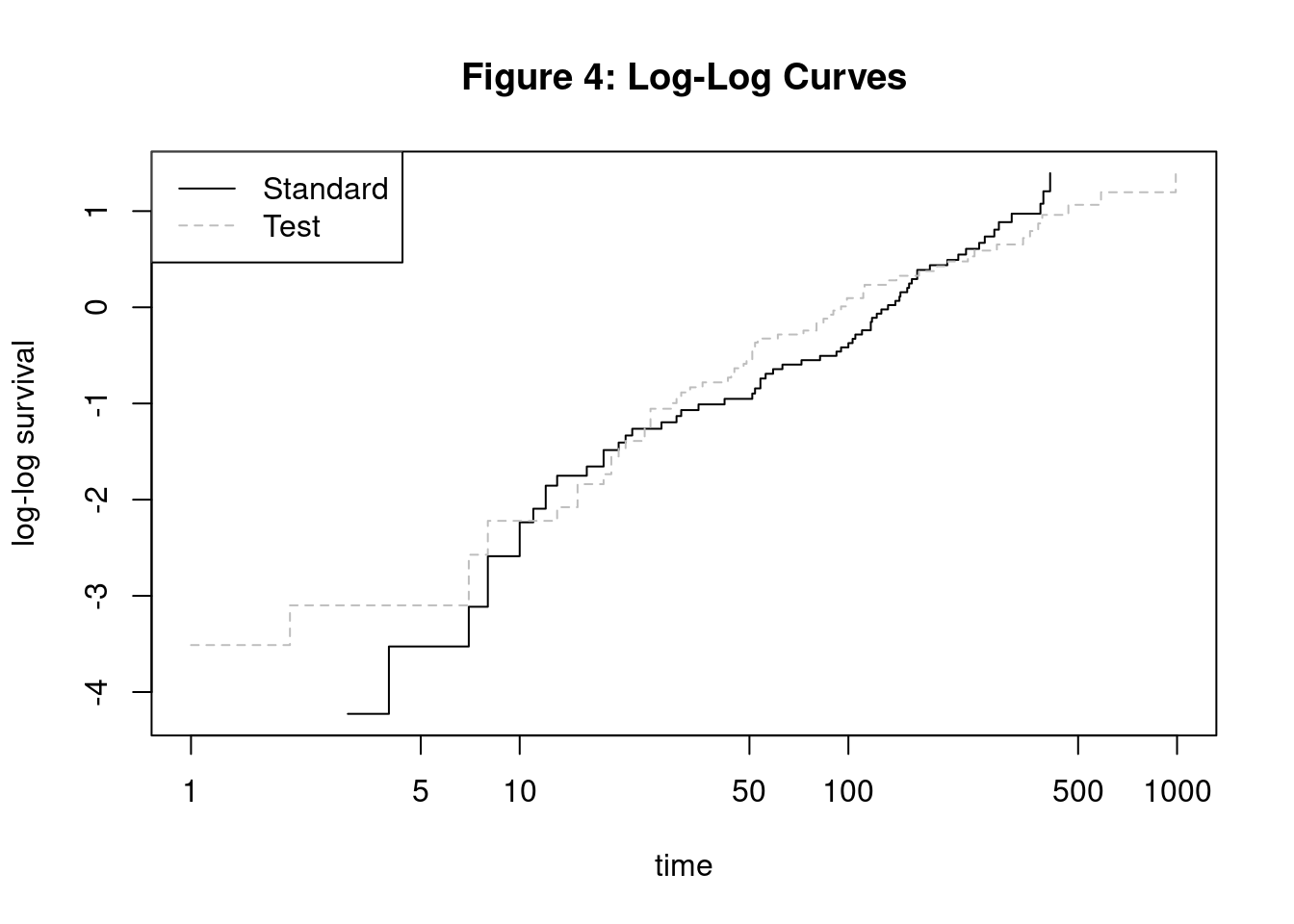
Log-Log Survival Curve
1. Residuals vs Time Plot
PH assumption is valid when coefficient is constant over time.
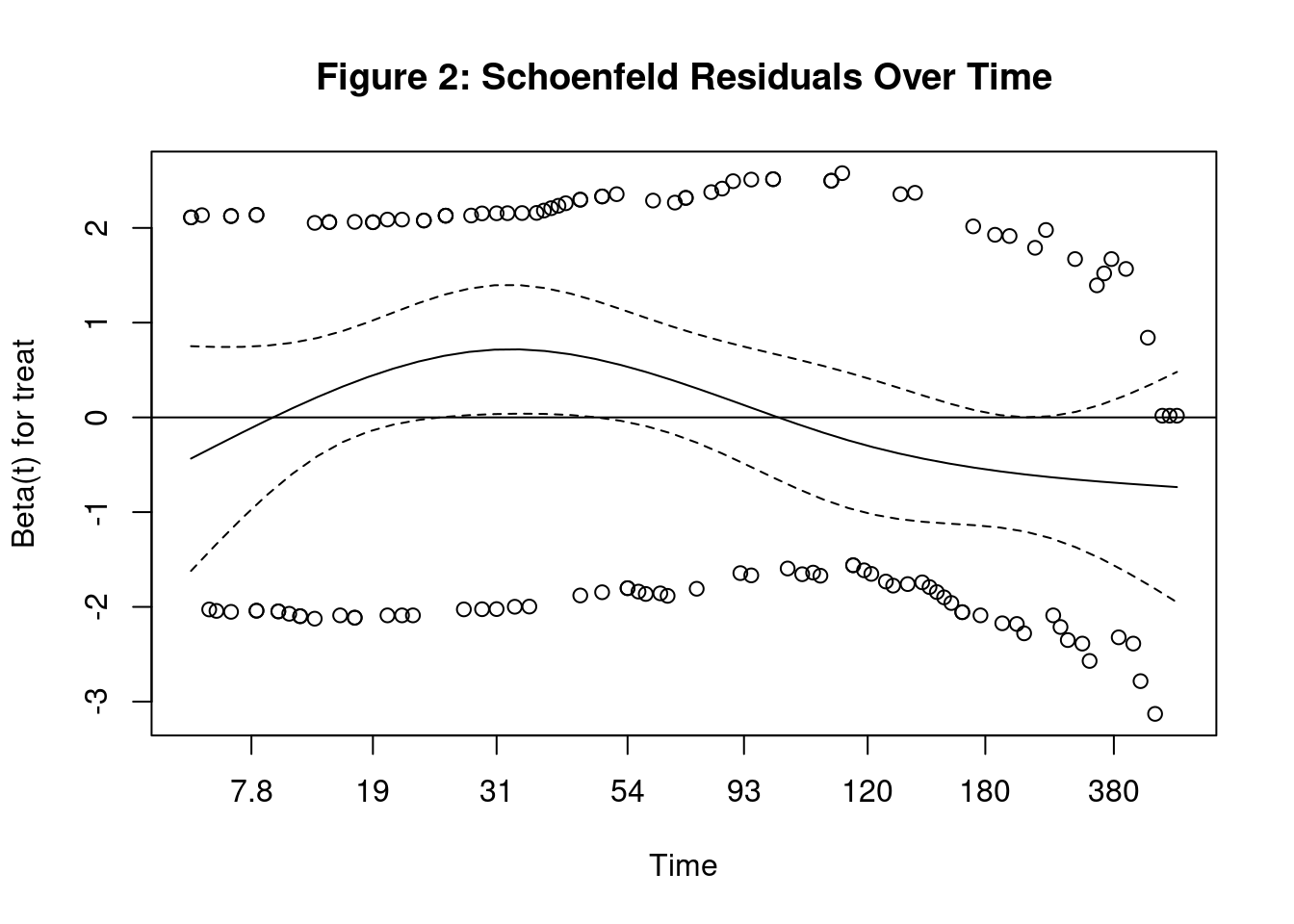
2. Statistical Test
Test if Schoenfeld residuals are correlated with time. The null hypothesis \(H_0\) is that the residuals are not correlated with time.
If p-value < \(\alpha\): 0.05, then PH assumption is violated.
3. Survival Curves
Multiple Variable Cox Model
Call:
coxph(formula = Y ~ treat + age + Karn + diag.time + cell + prior,
data = VA)
coef exp(coef) se(coef) z p
treat2 2.946e-01 1.343e+00 2.075e-01 1.419 0.15577
age -8.706e-03 9.913e-01 9.300e-03 -0.936 0.34920
Karn -3.282e-02 9.677e-01 5.508e-03 -5.958 2.55e-09
diag.time 8.132e-05 1.000e+00 9.136e-03 0.009 0.99290
cell2 8.616e-01 2.367e+00 2.753e-01 3.130 0.00175
cell3 1.196e+00 3.307e+00 3.009e-01 3.975 7.05e-05
cell4 4.013e-01 1.494e+00 2.827e-01 1.420 0.15574
prior10 7.159e-02 1.074e+00 2.323e-01 0.308 0.75794
Likelihood ratio test=62.1 on 8 df, p=1.799e-10
n= 137, number of events= 128 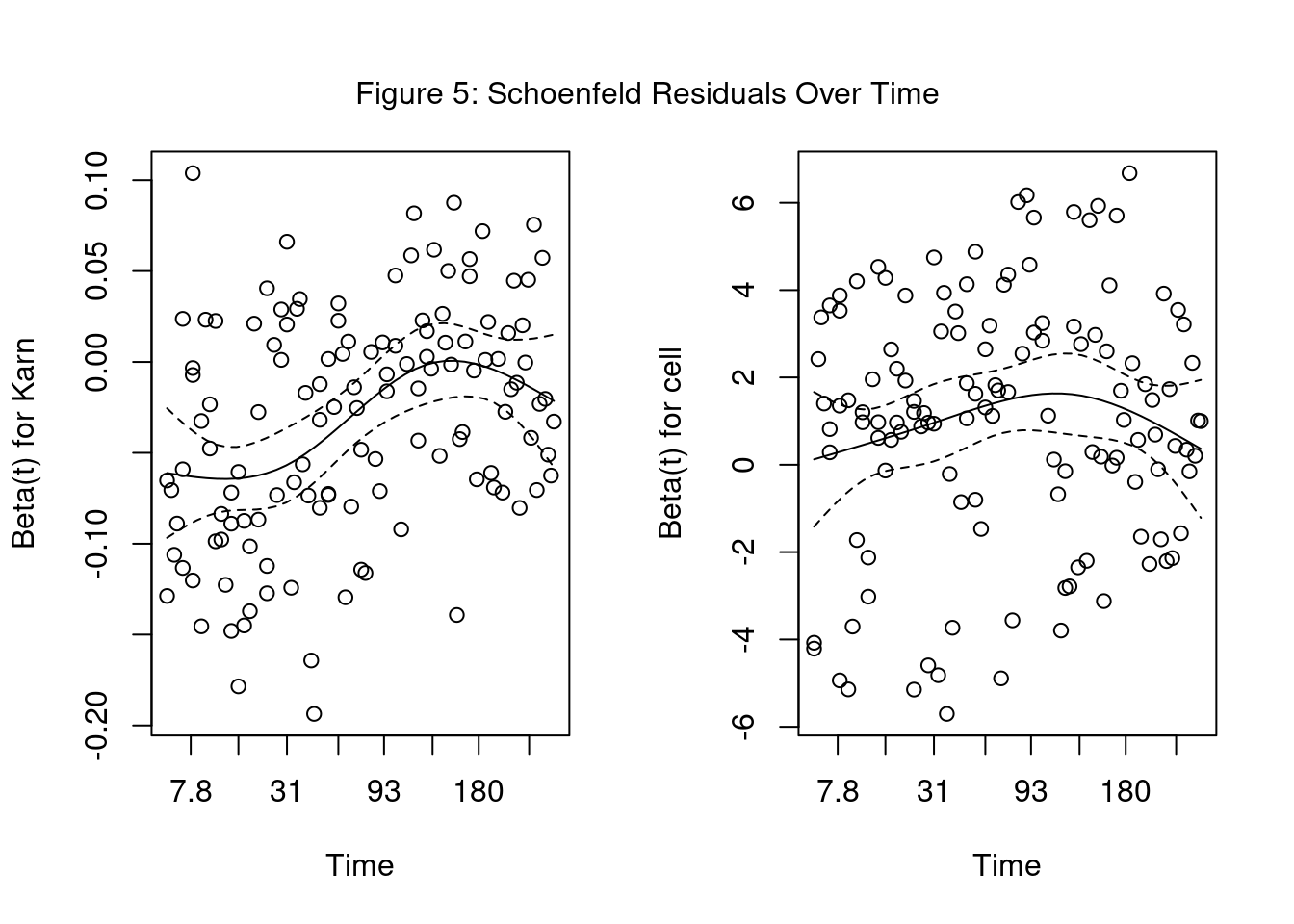
Options when PH assumption not met
Subset data based time
Start Cox PH model after certain time period (ie fit model for 180-day survivors)
Doesn’t consider individuals whose event occurred before 180 days
Stratified Cox model
Obtain separate models for each group (ie Cell Type)
Cannot compare HR across stratified groups
More Options when PH assumption not met
Extended Cox model
Specify function time-varying coefficients and/or time-dependent variables
Could be harder to interpret depending on function
Combination of 1, 2, or 3
1. Fitting Cox model for 180-day survivors
Only 27 individual left in study after 180 days.
Sample too small to make conclusions.
2. Stratified Cox model
Assumes a different baseline hazard function for each Cell Type.
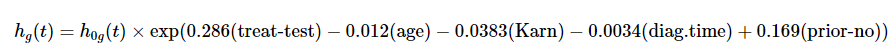
where \[g = 1, 2, 3, 4 \text{ (number of cell types)}\]
Call:
coxph(formula = Y ~ treat + age + Karn + diag.time + strata(cell) +
prior, data = VA)
coef exp(coef) se(coef) z p
treat2 0.285902 1.330962 0.210009 1.361 0.173
age -0.011821 0.988249 0.009846 -1.201 0.230
Karn -0.038262 0.962461 0.005932 -6.450 1.12e-10
diag.time -0.003439 0.996567 0.009075 -0.379 0.705
prior10 0.169069 1.184201 0.235667 0.717 0.473
Likelihood ratio test=44.27 on 5 df, p=2.042e-08
n= 137, number of events= 128 3. Extended Cox model
\[h_i(t)= h_0(t) \times exp(\beta_1(t)X_{1i} + ... + \beta_k(t)X_{ki}) \]Used to specify a function for the coefficients that change with time (linear, logarithmic, a step function, etc).
We used a step function for time intervals of 0-90, 90-180, and 180+ days.
#Creating tgroup column for each individual
VA.cp = survSplit(Surv(stime, status) ~ ., data=VA, cut=c(90,180), episode="tgroup")
# Output first 7 entries
VA.cp[1:7, c("age","Karn","cell","tstart", "stime", "status", "tgroup")] age Karn cell tstart stime status tgroup
1 69 60 1 0 72 1 1
2 64 70 1 0 90 0 1
3 64 70 1 90 180 0 2
4 64 70 1 180 411 1 3
5 38 60 1 0 90 0 1
6 38 60 1 90 180 0 2
7 38 60 1 180 228 1 3coxph(Surv(tstart, stime, status) ~ treat + age + diag.time + cell + prior + Karn:strata(tgroup),
data=VA.cp)Call:
coxph(formula = Surv(tstart, stime, status) ~ treat + age + diag.time +
cell + prior + Karn:strata(tgroup), data = VA.cp)
coef exp(coef) se(coef) z p
treat2 0.112060 1.118580 0.211447 0.530 0.596133
age -0.010939 0.989120 0.009358 -1.169 0.242412
diag.time -0.002008 0.997994 0.008781 -0.229 0.819087
cell2 0.959954 2.611577 0.285245 3.365 0.000764
cell3 1.144719 3.141558 0.306753 3.732 0.000190
cell4 0.345923 1.413293 0.284123 1.218 0.223411
prior10 0.084232 1.087881 0.234362 0.359 0.719290
Karn:strata(tgroup)tgroup=1 -0.048006 0.953128 0.006561 -7.317 2.53e-13
Karn:strata(tgroup)tgroup=2 0.007040 1.007065 0.013324 0.528 0.597241
Karn:strata(tgroup)tgroup=3 0.000932 1.000932 0.014981 0.062 0.950397
Likelihood ratio test=82.3 on 10 df, p=1.773e-13
n= 225, number of events= 128 4. Stratified-Extended (SE) Cox model
We can both stratify on cell type while using the extended model for the Karnofsky score.
coxph(Surv(tstart, stime, status) ~ treat + age + diag.time + strata(cell) + prior + Karn:strata(tgroup),
data=VA.cp)Call:
coxph(formula = Surv(tstart, stime, status) ~ treat + age + diag.time +
strata(cell) + prior + Karn:strata(tgroup), data = VA.cp)
coef exp(coef) se(coef) z p
treat2 0.160901 1.174569 0.215924 0.745 0.456
age -0.012389 0.987687 0.009893 -1.252 0.210
diag.time -0.004278 0.995731 0.008938 -0.479 0.632
prior10 0.164566 1.178882 0.237594 0.693 0.489
Karn:strata(tgroup)tgroup=1 -0.049032 0.952150 0.006818 -7.192 6.41e-13
Karn:strata(tgroup)tgroup=2 0.008194 1.008228 0.015464 0.530 0.596
Karn:strata(tgroup)tgroup=3 -0.019916 0.980281 0.020078 -0.992 0.321
Likelihood ratio test=57.35 on 7 df, p=5.104e-10
n= 225, number of events= 128 Reducing Model
We can use the Log-Likelihood value (\(L\)) to determine if there is a significant difference in nested models.
\[\chi_{LR}^2 = -2\ln L_\text{reduced-model} - (-2\ln L_\text{full-model})\]
If \(\chi_{LR}^2 > \chi_{df,\alpha}^2\) (ie p-value < 0.05), then significant difference in models
Analysis of Deviance Table
Cox model: response is Surv(tstart, stime, status)
Model 1: ~ treat + strata(cell) + Karn:strata(tgroup)
Model 2: ~ treat + age + diag.time + strata(cell) + prior + Karn:strata(tgroup)
loglik Chisq Df Pr(>|Chi|)
1 -311.09
2 -310.06 2.0444 3 0.5632Final SE Cox model
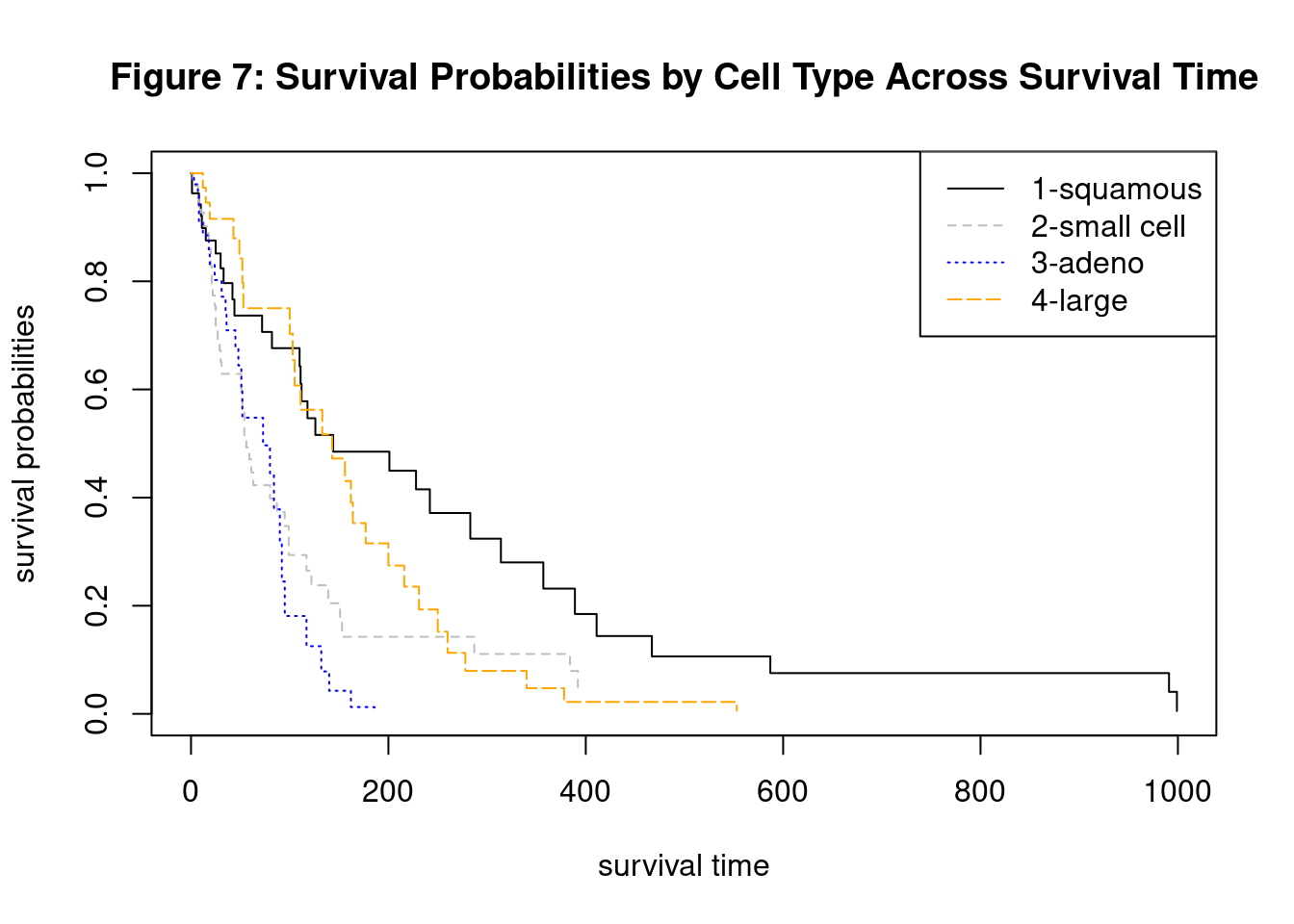
Simplified the model to treat + Karn + cell.
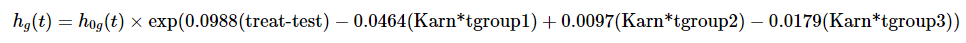
where \[g = 1, 2, 3, 4 \text{ (number of cell types)}\] \[tgroup1 = 1 \text{ if } 0 \le t < 90 \] \[tgroup2 = 1 \text{ if } 90 \le t < 180 \] \[tgroup3 = 1 \text{ if } t \ge 180\]
SE Cox model Summary
| Characteristic | HR1 | 95% CI1 | p-value |
|---|---|---|---|
| treat | |||
| 1 | — | — | |
| 2 | 1.10 | 0.74, 1.66 | 0.6 |
| Karn * strata(tgroup) | |||
| Karn * tgroup=1 | 0.95 | 0.94, 0.97 | <0.001 |
| Karn * tgroup=2 | 1.01 | 0.98, 1.04 | 0.5 |
| Karn * tgroup=3 | 0.98 | 0.95, 1.02 | 0.4 |
| Log-likelihood | -311 | ||
| c-index | 0.705 | ||
| 1 HR = Hazard Ratio, CI = Confidence Interval | |||
SE Cox model Interpretations
- Treatment effect was not found to be significant, with a p value of 0.6, in final Statified-Extended Cox model
Karnofsky’s score was found to be significant but only in the first 90 days, with a p value <0.001
In the first 90 days, for a 10-point decrease in Karnofsky score the probability of dying for a patient increase by 57%, all else constant
Karnofsky score is measured at the time of entry to the trial
Concluding Remarks
Cox Proportional Hazards Model is a useful tool in survival analysis
Regarding the case-study results…
It is due to the CPH’s robust nature and being able to handle censored data, which leads it to be so popular in survival analysis, as seen in our case-study